Dunn Laboratory

Dunn Lab
Department of Neurosurgery
Brain Tumor Immunology and Immunotherapy Program
Massachusetts General Hospital
Harvard Medical School

Our location
Our lab is located in the Simches Research Center
on the main Massachusetts General Hospital ( MGH )campus
185 Cambridge St, Boston, MA 02114

Our Mission
Using immunology and genomics tools, we study how the immune response to brain tumors happens. We use established preclinical models, new preclinical models we have developed, and patient-derived tissue and cells to address what T cells recognize and how antigen is presented. These studies are complemented by clinical trial efforts. Our ultimate goal is to better understand the diseases of our patients in order to develop improved therapies.
Select Publications

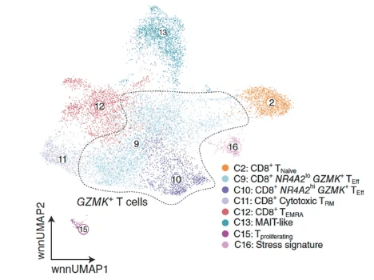
Glioblastoma-Infiltrating CD8+ T Cells Are Predominantly a Clonally Expanded GZMK+ Effector Population
Cancer Discov. 2024 Jun 3;14(6):1106-1131
Recent clinical trials have highlighted the limited efficacyof T cell-based immunotherapy in patients with glioblastoma (GBM). To better understand the characteristics of tumor-infiltrating lymphocytes (TIL) in GBM, we performed cellular indexing of transcriptomes and epitopes by sequencing and single-cell RNA sequencing with paired V(D)J sequencing, respectively, on TILs from two cohorts of patients totaling 15 patients with high-grade glioma, including GBM or astrocytoma, IDH-mutant, grade 4 (G4A). Analysis of the CD8+ TIL landscape reveals an enrichment of clonally expanded GZMK+ effector T cells in the tumor compared with matched blood, which was validated at the protein level. Furthermore, integration with other cancer types highlights the lack of a canonically exhausted CD8+ T-cell population in GBM TIL. These data suggest that GZMK+ effector T cells represent an important T-cell subset within the GBM microenvironment and may harbor potential therapeutic implications.
Read more here: https:://pubmed.ncbi.nlm.nih.gov/38416133/

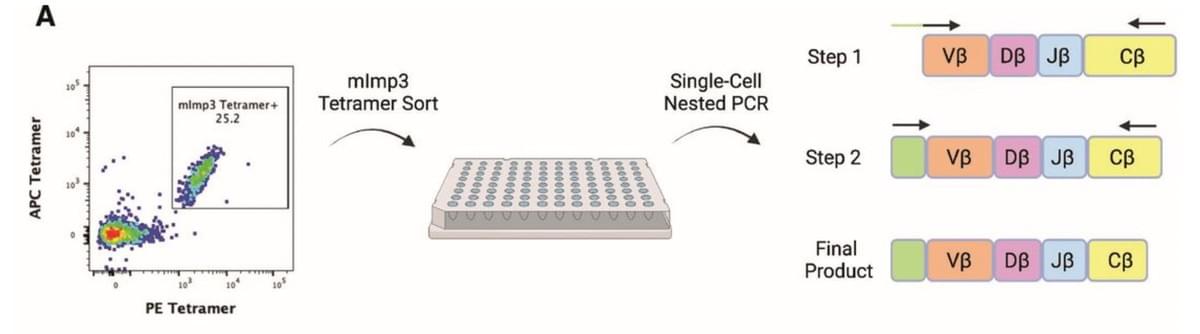
TCR-engineered adoptive cell therapy effectively treats intracranial murine glioblastoma
J Immunother Cancer. 2023 Feb;11(2):e006121.
We employed single-cell PCR to isolate a TCR specific for the Imp3D81N neoantigen (mImp3) previously identified within the murine glioblastoma model GL261. This TCR was used to generate the Mutant Imp3-Specific TCR TransgenIC (MISTIC) mouse in which all CD8 T cells are specific for mImp3. The therapeutic efficacy of neoantigen-specific T cells was assessed through a model of cellular therapy consisting of the transfer of activated MISTIC T cells and interleukin 2 into lymphodepleted tumor-bearing mice. In a model of adoptive cellular therapy, the infusion of activated MISTIC T cells resulted in rapid intratumoral infiltration and profound antitumor effects with long-term cures in a majority of GL261-bearing mice. The subset of mice that did not respond to the adoptive cell therapy showed evidence of retained neoantigen expression but intratumoral MISTIC T cell dysfunction. The efficacy of MISTIC T cell therapy was lost in mice bearing a tumor with heterogeneous mImp3 expression, showcasing the barriers to targeted therapy in polyclonal human tumors.
Read more here: https://pubmed.ncbi.nlm.nih.gov/36808076/

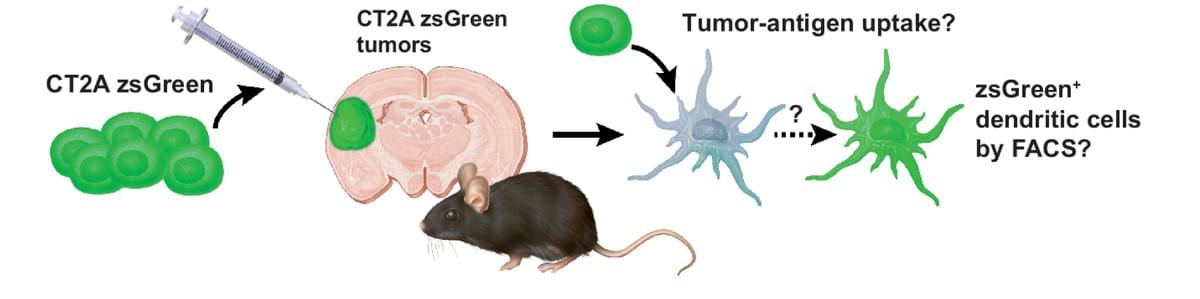
The Conventional Dendritic Cell 1 Subset Primes CD8+ T Cells and Traffics Tumor Antigen to Drive Antitumor Immunity in the Brain
Cancer Immunol Res. 2023 Jan 3;11(1):20-37. doi: 10.1158/2326-6066.CIR-22-0098.
The central nervous system (CNS) antigen-presenting cell (APC) that primes antitumor CD8+ T-cell responses remains undefined. Elsewhere in the body, the conventional dendritic cell 1 (cDC1) performs this role. However, steady-state brain parenchyma cDC1 are extremely rare; cDCs localize to the choroid plexus and dura. Thus, whether the cDC1 play a function in presenting antigen derived from parenchymal sources in the tumor setting remains unknown. Using preclinical glioblastoma (GBM) models and cDC1-deficient mice, we explored the presently unknown role of cDC1 in CNS antitumor immunity. We determined that, in addition to infiltrating the brain tumor parenchyma itself, cDC1 prime neoantigen-specific CD8+ T cells against brain tumors and mediate checkpoint blockade-induced survival benefit. We observed that cDC, including cDC1, isolated from the tumor, the dura, and the CNS-draining cervical lymph nodes harbored a traceable fluorescent tumor antigen. In patient samples, we observed several APC subsets (including the CD141+ cDC1 equivalent) infiltrating glioblastomas, meningiomas, and dura. In these same APC subsets, we identified a tumor-specific fluorescent metabolite of 5-aminolevulinic acid, which fluorescently labeled tumor cells during fluorescence-guided GBM resection. Together, these data elucidate the specialized behavior of cDC1 and suggest that cDC1 play a significant role in CNS antitumor immunity.
Read more here: https://pubmed.ncbi.nlm.nih.gov/36409838/

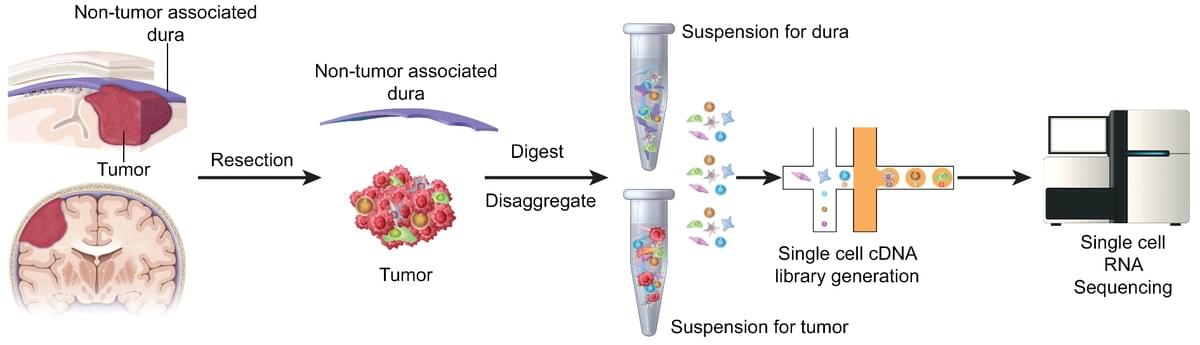
Single-cell profiling of human dura and meningioma reveals cellular meningeal landscape and insights into meningioma immune response
Genome Med. 2022 May 10;14(1):49. doi: 10.1186/s13073-022-01051-9
We use single-cell RNA sequencing to perform the first characterization of both non-tumor-associated human dura and primary meningioma samples. First, we reveal a complex immune microenvironment in human dura that is transcriptionally distinct from that of meningioma. In addition, we characterize a functionally diverse and heterogenous landscape of non-immune cells including endothelial cells and fibroblasts. Through imaging mass cytometry, we highlight the spatial relationship among immune cell types and vasculature in non-tumor-associated dura. Utilizing T cell receptor sequencing, we show significant TCR overlap between matched dura and meningioma samples. Finally, we report copy number variant heterogeneity within our meningioma samples. Our comprehensive investigation of both the immune and non-immune cellular landscapes of human dura and meningioma at single-cell resolution builds upon previously published data in murine models and provides new insight into previously uncharacterized roles of human dura.
Read more here: https://pubmed.ncbi.nlm.nih.gov/35534852/

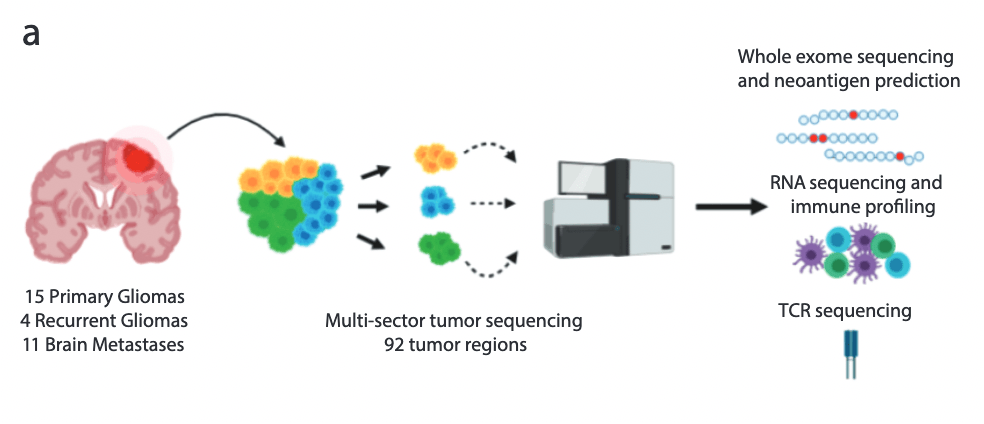
Characterization of the Genomic and Immunological Diversity of Malignant Brain Tumors Through Multi-Sector Analysis
Cancer Discov. 2021 Oct 5:candisc.0291.2021
Despite some success in secondary brain metastases, targeted or immune-based therapies have shown limited efficacy against primary brain malignancies such as glioblastoma (GBM). While the intratumoral heterogeneity of GBM is implicated in treatment resistance, it remains unclear whether this diversity is observed within brain metastases and to what extent cancer-cell intrinsic heterogeneity sculpts the local immune microenvironment. Here, we profiled the immunogenomic state of 93 spatially distinct regions from 30 malignant brain tumors through whole exome, RNA, and TCR-sequencing. Our analyses identified differences between primary and secondary malignancies with gliomas displaying more spatial heterogeneity at the genomic and neoantigen level. Additionally, this spatial diversity was recapitulated in the distribution of T cell clones where some gliomas harbored highly expanded but spatially restricted clonotypes. This study defines the immunogenomic landscape across a cohort of malignant brain tumors and contains implications for the design of targeted and immune-based therapies against intracranial malignancies.
Read more here: https://pubmed.ncbi.nlm.nih.gov/34610950/

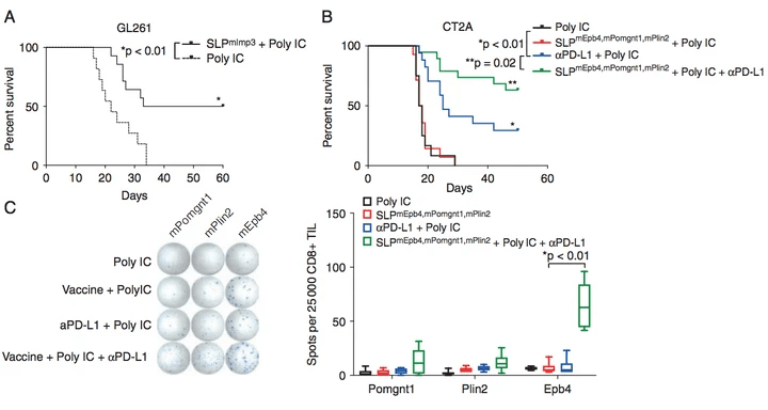
Treatment of an Aggressive Orthotopic Murine Glioblastoma Model with Combination Checkpoint Blockade and a Multivalent Neoantigen Vaccine
Neuro-Oncol. 2020 Mar 5;noaa050
We characterized the microenvironment and neoantigen landscape of the aggressive CT2A GBM model in order to develop a platform to test combination checkpoint blockade and neoantigen vaccination. Whole exome DNA and RNA sequencing of the CT2A murine GBM was employed to identify expressed, somatic mutations for vaccine development. Survival analysis showed that therapeutic neoantigen vaccination with Epb4H471L, Pomgnt1R497L, and Plin2G332R, in combination with αPD-L1 treatment was superior to αPD-L1 alone. These observations provide important preclinical correlates for GBM immunotherapy trials and support further investigation into the effects of multi-modal immunotherapeutic interventions on anti-glioma immunity
Read more here: https://www.ncbi.nlm.nih.gov/pubmed/32133512

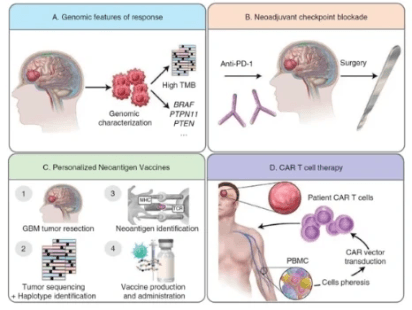
Emerging Immunotherapies for Malignant Glioma: From Immunogenomics to Cell Therapy
Neuro Oncol. 2020 Jul 2: noaa154
As immunotherapy assumes a central role in the management of many cancers, ongoing work is directed at understanding whether immune-based treatments will be successful in patients with glioblastoma (GBM). Despite several large studies conducted in the last several years, there remain no FDA-approved immunotherapies in this patient population. Nevertheless, there are a range of exciting new approaches being applied to GBM, all of which may not only allow us to develop new treatments but also help us understand fundamental features of the immune response in the central nervous system. In this review, we summarize new developments in the application of immune checkpoint blockade, from biomarker-driven patient selection to the timing of treatment. Moreover, we summarize novel work in personalized immune-oncology by reviewing work in cancer immunogenomics–driven neoantigen vaccine studies. Finally, we discuss cell therapy efforts by reviewing the current state of chimeric antigen receptor T-cell therapy.
Read more here: https://www.ncbi.nlm.nih.gov/pubmed/32615600

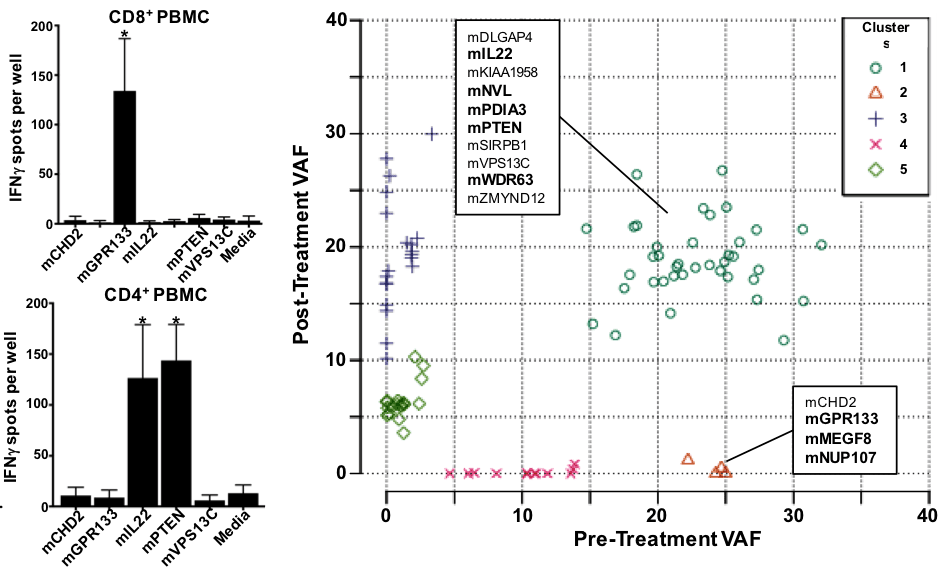
Detection of Neoantigen-specific T Cells Following a Personalized Vaccine in a Patient with Glioblastoma
Oncoimmunology, 2019 Jan 25;8(4):e1561106
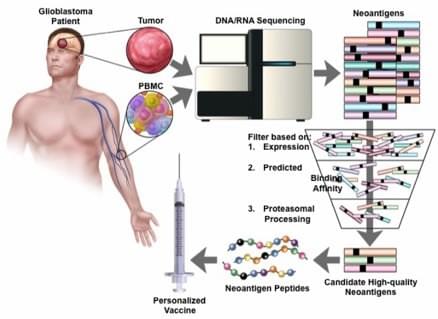
Neoantigens represent promising targets for personalized cancer vaccine strategies. However, the feasibility of this approach in lower mutational burden tumors like glioblastoma (GBM) remains unknown. We report the application a cancer immunogenomics pipeline to identify candidate neoantigens and guide screening for neoantigen-specific T cell responses in a patient with GBM treated with a personalized synthetic long peptide vaccine. Following vaccination, reactivity to 3 HLA class I- and 5 HLA class II-restricted candidate neoantigens were detected. These data demonstrate the feasibility and translational potential of a therapeutic neoantigen-based vaccine approach in patients with primary CNS tumors.
Read more here: https://www.ncbi.nlm.nih.gov/pubmed/30906654

Targeting Neoantigens in Glioblastoma: An Overview of Cancer Immunogenomics and Translational Implications
Neurosurgery. 2017 Sep 1;64(CN_suppl_1):165-176
Recent interest has been focused on the identification of tumor-specific mutations, termed neoantigens, which can serve as immunodominant targets for antitumor immune effector cells to maximize “on-tumor” effect and minimize “off-tumor” toxicities. In this review, we discuss: (1) the current perspective on CNS immunosurveillance, (2) the process of neoantigen identification focusing on the cancer immunogenomics approach, and (3) how this strategy can be used to target GBM specifically.
Read more here: https://www.ncbi.nlm.nih.gov/pubmed/28899059

Immunogenomics of Hypermutated Glioblastoma: A Patient with Germline POLE Deficiency Treated with Checkpoint Blockade Immunotherapy
Cancer Discov. 2016 Nov;6(11):1230-1236.
We present the case of a patient with a left frontal glioblastoma with primitive neuroectodermal tumor features and hypermutated genotype in the setting of a POLE germline alteration. Using whole-exome DNA sequencing and clonal analysis, we report changes in the subclonal architecture throughout treatment. Furthermore, a persistently high neoantigen load was observed within all tumors. Interestingly, following initiation of pembrolizumab, brisk lymphocyte infiltration was observed in the subsequently resected metastatic spinal lesion and an objective radiographic response was noted in a progressive intracranial lesion, suggestive of active central nervous system (CNS) immunosurveillance following checkpoint blockade therapy.
Read more here: https://www.ncbi.nlm.nih.gov/pubmed/27683556

Endogenous Neoantigen-Specific CD8 T Cells Identified in Two Glioblastoma Models Using a Cancer Immunogenomics Approach
Cancer Immunol Res. 2016 Dec;4(12):1007-1015
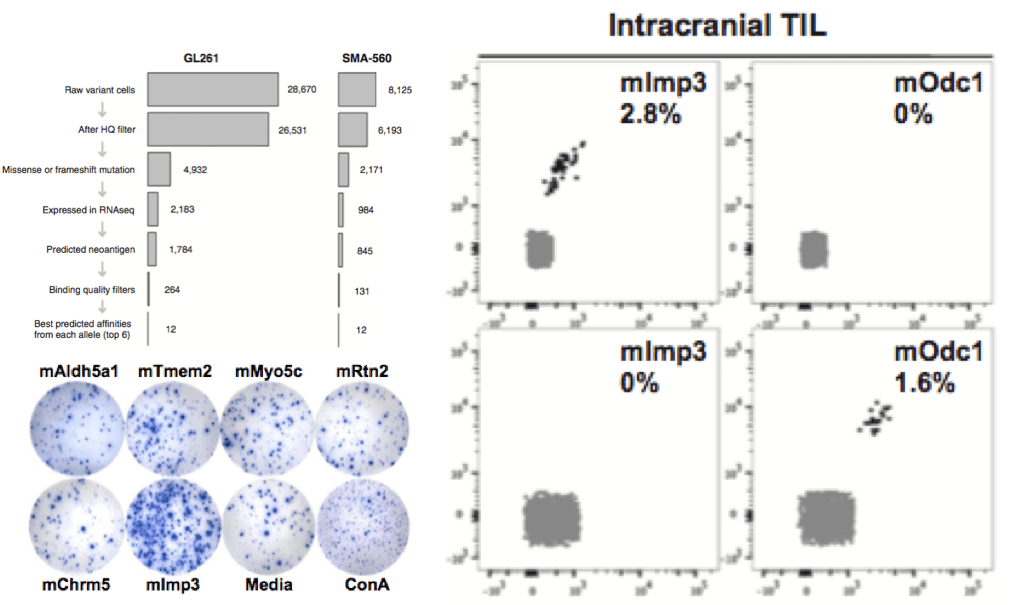
We applied a cancer immunogenomics approach to identify tumor-specific "neoantigens" in the C57BL/6-derived GL261 and VM/Dk-derived SMA-560 tumor models. Following DNA whole-exome and RNA sequencing, high-affinity candidate neoepitopes were predicted and screened for immunogenicity by ELISPOT and tetramer analyses. We confirmed H-2Db-restricted endogenous tumor-specific neoantigens that are functionally immunogenic. By establishing the immunogenicities of predicted high-affinity neoepitopes in these models, we extend the immunogenomics-based neoantigen discovery pipeline to glioblastoma models and provide a tractable system to further study the mechanism of action of T cell-activating immunotherapeutic approaches in preclinical models of glioblastoma
Read more here: https://www.ncbi.nlm.nih.gov/pubmed/27799140
Funding

© 2025